In the intricate world of microbiome research, a groundbreaking method has emerged that could revolutionize how scientists study host-microbe interactions, particularly in samples where host DNA dominates. This innovation, known as 2bRAD-M, is poised to make waves not only in human health but also in the agriculture sector, offering a more efficient and practical approach to microbiome analysis.
The challenge of characterizing human microbiota in host-dominated samples has long been a hurdle for researchers. Traditional methods often require host depletion, a process that can lead to DNA loss and necessitates immediate sample processing. Enter 2bRAD-M, a reduced metagenomic sequencing method that bypasses the need for host depletion, as detailed in a recent study published in *npj Biofilms and Microbiomes*.
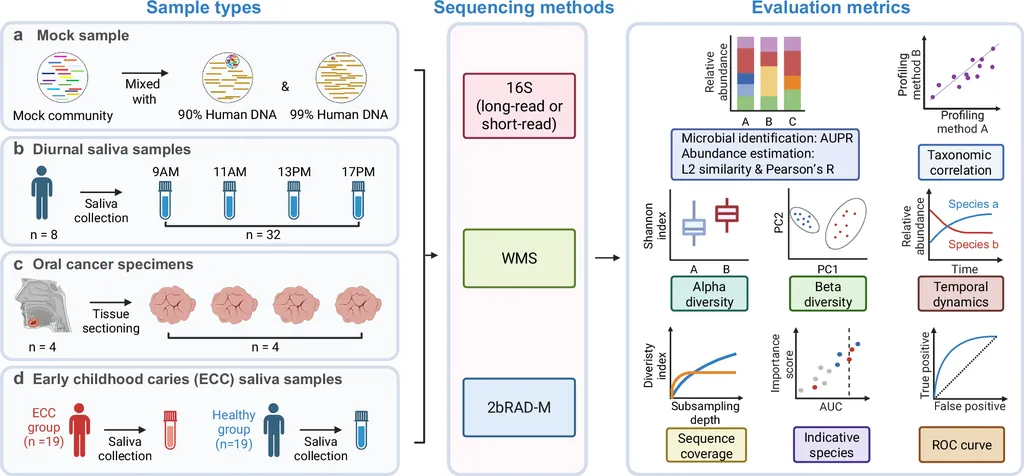
Led by Yuesong Jiang from the Faculty of Dentistry at The University of Hong Kong, the research team validated 2bRAD-M on mock samples containing over 90% human DNA. The results were impressive, with the method achieving over 93% in AUPR and L2 similarity. “This method provides a high-resolution microbial profile without the need for host depletion, which is a significant advancement in the field,” Jiang explained.
The practical implications of this technology are vast. In agriculture, understanding the microbiome of plants and animals is crucial for improving crop yields, enhancing animal health, and developing sustainable farming practices. Traditional methods of microbiome analysis can be time-consuming and costly, often requiring extensive sample preparation and processing. 2bRAD-M offers a more streamlined approach, potentially reducing the sequencing effort by up to 90% while still capturing detailed microbial profiles.
For instance, in the study, 2bRAD-M was used to analyze saliva and oral cancer samples, closely matching whole metagenomic sequencing (WMS) profiles. In saliva samples, it captured diurnal and host-specific patterns with only 5–10% of the sequencing effort. This efficiency could translate to significant cost savings and faster turnaround times in agricultural research, allowing scientists to study the microbiomes of crops and livestock more effectively.
Moreover, the method’s ability to distinguish between healthy and diseased states, as demonstrated in an early childhood caries (ECC) study, highlights its potential for diagnostic applications. In agriculture, similar principles could be applied to identify microbial markers associated with plant health or disease, enabling early intervention and prevention strategies.
The commercial impact of 2bRAD-M could be profound. By providing a more efficient and cost-effective means of microbiome analysis, it could accelerate research and development in the agriculture sector, leading to innovative solutions for sustainable farming and food security. As Jiang noted, “This method offers a practical and efficient solution for microbiome research, which can have far-reaching implications across various fields, including agriculture.”
In the broader context, the development of 2bRAD-M underscores the importance of interdisciplinary research. By bridging the gap between human health and agricultural science, this method exemplifies how advancements in one field can catalyze progress in another. As we look to the future, the potential applications of 2bRAD-M are limited only by our imagination, promising to shape the next generation of microbiome research and its impact on global agriculture.